This document is also available in PDF format and in other languages: español, français.

Colophon
Suggested citation
Chapman AD (2020) Current Best Practices for Generalizing Sensitive Species Occurrence Data. Copenhagen: GBIF Secretariat. https://doi.org/10.15468/doc-5jp4-5g10.
Licence
The document Current Best Practices for Generalizing Sensitive Species Occurrence Data is licensed under Creative Commons Attribution-ShareAlike 4.0 Unported License.
Document control
Version 1.0, November 2020
Based on the earlier publication by Chapman AD & Grafton O (2008) Guide to Best Practices for Generalising Sensitive Species-Occurrence Data. Copenhagen: GBIF Secretariat. https://doi.org/10.15468/doc-b02j-gt10.
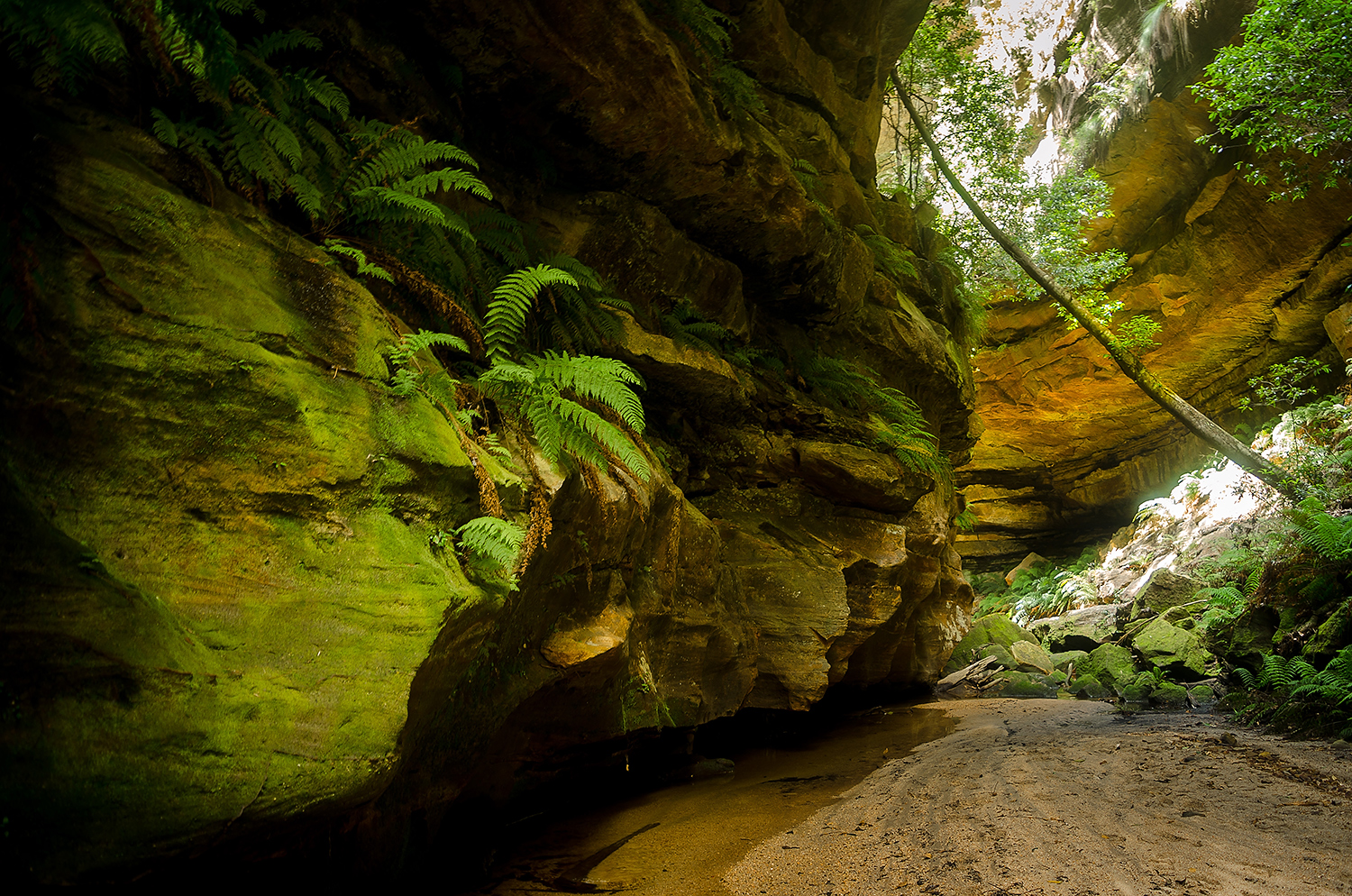
Cover image
Beneath the surface, Wollemi National Park, Australia. Photo 2015 Vern via Flickr, licensed under CC BY-NC-ND 2.0.
Introduction
The unprotected distribution of Sensitive Primary Species Occurrence Data (for example the exact localities of rare, endangered or commercially valuable taxa) was a concern of GBIF – the Global Biodiversity Information Facility – from its beginning. The GBIF Secretariat has a vested interest in making data available via its portals, but at the same time respecting the wishes of data providers to restrict information on sensitive taxa. In early 2006, GBIF initiated a process to address this issue, especially in relation to data to be shared through the GBIF network and made visible through GBIF.org and other data aggregating initiatives.
This resulted in the Guide to Best Practices for Generalising Sensitive Primary Species Occurrence Data. That document relied heavily on the results of an online survey conducted through Survey Monkey and subsequent workshops whose reports were originally made available on the GBIF website (Chapman 2006).
A final report on Dealing with Sensitive Primary Species Occurrence Data was developed following these processes and discussions, and was presented to GBIF in April 2007 (Chapman 2007). This report made a number of recommendations, and many of these have been included in this document.
The final step in that process was to develop a Guide to Best Practices for Primary Species Occurrence Data. That document was proposed as an overriding guideline for institutions, data providers and GBIF Nodes to use to develop their own in-house guidelines. Organizations and institutions were encouraged to produce their own internal documents that incorporated the practices outlined in the Guide and related documents such as the Guide to Best Practices for Georeferencing (Chapman and Wieczorek 2006) and incorporate them into their own working environment. Unfortunately, not as many institutions have taken up the challenge and produced their own internal documents as we had hoped. Two key agencies that have done so, however, are SANBI in South Africa (SANBI 2010) and the Atlas of Living Australia (Tann and Flemons 2009, ALA 2018a) (see Implementations).
It is also important to understand the possible impact that approaches for restricting sensitive data may have on biodiversity science and, while restricting the availability or resolution of certain data, not overly restricting the uses to which the data may be put. For that reason, a set of principles are elucidated below. Key among these is the need to make biodiversity information freely available wherever possible, in the interests of science, the environment and biodiversity itself.
| Words hyperlinked in the text refer to terms that are included in the Glossary; citations link to sources where they are available online and to the References where they are not. |
Objectives
This document aims to provide best practice (or best current practice) for dealing with sensitive primary species occurrence data, and provide guidance on how to make as much data available without at the same time opening up the species to harm because data has been placed in the public domain.
It is now more than a decade since the first Guide was published, and this new publication is designed to bring those practices up to date and to incorporate the experiences gained by institutions that have implemented the Guide in whole or in part.
Audience
This work is designed for those who need, or want to know how they can best make as much data available on sensitive taxa as possible without that published data leading to harm to the species. This document is also for individuals or organizations faced with developing a policy on dealing with sensitive primary species occurrence data and writing in-house documentation consistent with current best practice.
Above all, this document will help end users of the data to understand the implications of trying to use records that may have been generalized to protect sensitive species, and how to understand the meaning of generalization at different precisions.
Scope
The term “best practice” generally refers to the best possible way of doing something. It is commonly used in the fields of business management, software engineering, and medicine, and increasingly in government. The term “current best practice” (or best current practice) is more specific in that it indicates the possibility for future developments and better practice. Due to the immaturity of this topic, this publication generally refers to “current” best practice and is sure to mature over time as more institutions adopt and adapt the principles outlined herein.
Two issues that this and the previous document have not covered are the issues of the privacy of living individuals and the development of data sharing and data licensing agreements. Both of these issues have legal implications and vary considerably from jurisdiction to jurisdiction. These issues have been covered in detail by others (e.g. Corti et al. 2000, Parry and Mauthner 2004, GBIF 2017, GBIF 2019, ALA 2018b, OEH 2019b).
Ethics and biodiversity is a topic that has received little coverage, although for hundreds of years biologists have followed implied ethics in their work. The management of data on sensitive data requires considerable ethical practice and in many cases, a lot of trust and collaboration. Often amateur biologists and citizen scientists are aware of the locations of sensitive taxa, and it is up to the biologists to work with these groups to ensure the continued survival of the species. This won’t always be possible as there will always be rogues, but collaboration can work. For example, working with amateur groups to ethically bring rare plants into cultivation so that there is less pressure on the wild populations.
Archaeology has been grappling with these issues for a long time. As Alison Wylie (1996) explains,
the foundation of the Society for American Archaeology (SAA) was partially motivated by the desire of academics to distance themselves from amateur archaeologists through the establishment of ethical codes of conduct. One of the main concerns in the field has long been the practice of looting historical sites and the threat this poses to both cultural heritage and future archaeological work.
Nineteen years later, the archaeologists continue to grapple as pointed out by Frank et al. (2015) in discussing the results of 62 interviews of archaeologists and zoologists conducted by them. They concluded that “Researchers would generally prefer to restrict access to their data from the general public but maintain open data for colleagues”, while realizing the extreme difficulties in managing such a process.
This publication is largely focused on sensitive primary species occurrence data (e.g. museum and herbarium specimen data, observations, images, tracking data), but does have impacts on related data such as population sizes, numbers and viability, habitats and ecosystems, biogeography, traits, bycatch, biosecurity, etc.
This document only looks at taxon sensitivity and not sensitivity related to issues such as land or personal privacy. Different national and state jurisdictions have diverse legislation with regard to both land and personal privacy. Bearing that in mind, however, I would recommend following the same principles of generalization recommended in this publication wherever possible, rather than other methods such as randomization.
1. Principles
| Biodiversity information should be made freely available to be shared globally to enable their use for not-for-profit decision making, education, research and other public benefit purposes. Making the full detail of biodiversity information available should reduce the risk of damage to the environment and help safeguard a sustainable future. Where release will have the opposite effect, access to the full detail may need to be controlled. |
Below are a set of high-level principles related to the sharing of data generally and the sharing of sensitive data in particular.
-
The management of sensitive data is integral to ethical data management.
-
Wherever possible, environmental information should be freely available to all. Generally this benefits the environment by increasing awareness, enabling better decision-making and reducing risk of damage.
-
Public release of information can sometimes result in environmental harm. In such cases availability of information may need to be controlled; although the presumption remains in favour of release and any restrictions should be assessed and reviewed rigorously.
-
All data regarded as being sensitive should include a date for review of their sensitivity status, along with documented reasons for the sensitivity status. The date for review may be short or long depending on the nature of the sensitivity.
| Whenever a data provider receives an application for enhanced access to restricted data, they should avoid assuming continued sensitivity and use it as an opportunity to revisit the determination. |
-
If the data is to be restricted for distribution, then this should only be done to a copy of the data at the time of their distribution. Data should never be altered, falsified or deleted from the stored record.
-
Documentation is essential for many reasons, and where data have been restricted or generalized, it is important that the reason(s) for the categorization is recorded as metadata that remains with the record.
-
Where data is restricted or generalized for distribution (such as the name of a collector, textual locality information, etc.), this should be documented by replacing with appropriate wording − the field should not be left blank or null.
-
There are extremely strong reasons not to restrict data on related collections (e.g. collector’s numbers in sequence, collector’s name, etc.) because of the restrictions this places on data quality and data validation procedures, etc.
-
Users of sensitive data should comply with any and all restrictions of access that the data provider has placed on the data. If granted enhanced access to restricted information, users must not compromise or otherwise infringe the confidentiality of such information.
-
Data providers should respect the needs of data users to have access to data and documentation in order to determine the ‘fitness for use’ of the data and to ensure that analyses are robust and not misleading.
2. Determining sensitivity
As a first step, information holders need to identify any data which are regarded as ‘sensitive’. Sensitive information is any, which if released to the public, would result in an ‘adverse effect’ on the taxon or attribute in question or to a living individual. A number of factors need to be taken into account when determining sensitivity, including the type and level of threat, vulnerability of the taxon or attribute, type of information and whether it is already publicly available. Determining these factors leads us to a criteria-based approach.
Information cannot be considered sensitive if it is readily available through other sources or if it is not unique. This principle has been identified in a number of sensitive data policies (AMEC Earth and Environmental 2010, Australian Government 2016).
It would appear that herbaria are more inclined to restrict their data than mammal or insect collections (Chapman 2006). Perhaps this is because plants don’t move and the exact location of a collection is likely to lead one to an actual plant on the ground, whereas mammals and insects tend to move around. One entomologist commented that professional collectors and amateur groups often know more than the scientists about the location of rare species. However, there are categories of animals where the exact locations were thought to be sensitive and included bat roosting and maternity sites, nesting sites of falcons, and the location of various lizards, tortoise, butterfly species and large mammals. With plants, there is also a strong leaning towards not making information available for plants likely to be collected (pirated) such as cacti in Arizona, orchids and cycads. The protection of sensitive fossil sites was also identified. One unfortunate aspect is the susceptibility of a small number of taxa in a group (such as a few charismatic cacti, or orchids, etc.). This can often mean that all taxa in that group are then regarded as sensitive and the data on them restricted, even though many of those taxa are not themselves sensitive or susceptible to harmful acts.
On the other hand, some institutions have found benefit in working with the general public to gather information and to protect rare taxa, using the public and special interest groups to survey existing locations and to help locate new locations. There are good examples with birds, lizards, frogs, butterflies and various plant species (including orchids) in a number of countries. Several people have raised the issue of the balance between protecting taxa through knowledge of where they occur as opposed to protection through restricting knowledge of their occurrence at a location. This is very taxon (and maybe region) specific and certain taxa may be in greater danger due to inadvertent destruction through lack of knowledge than through deliberate collection and destruction through knowledge of locations. For this reason, a list of sensitive taxa should be quite different to a list of rare or threatened taxa, although there is likely to be considerable overlap between the two. It should be noted, also, that what is sensitive today, may not be sensitive tomorrow and vice versa, and this should lead to review on a periodic basis to determine whether the context has changed over time (AMEC Earth and Environmental 2010).
As noted in an article in Science (Stuart et al. 2006), three newly discovered amphibian and reptile species rapidly appeared in commercial trade shortly after their descriptions in the scientific literature. This is an issue of concern to biologists and especially to taxonomists (Guterman 2006) – how much information should be released in publication when describing a new taxon. There are a number of examples in coral-reef fish where a new species has appeared in the commercial trade soon after it is scientifically described.
A few examples with which I have had direct experience include Centropyge boylei, Centropyge narcosis [and] Belonoperce pylei … among a number of others.
Often in these and other cases, the existence of the new species is brought to the attention of the scientific community ‘by’ the commercial (aquarium) trade; rather than the other way around. Thus, it is usually not considered so much of a ‘problem’, but rather a sort of ‘symbiotic’ relationship between the commercial trade and the taxonomists. Moreover, in such cases in reef fishes, the species has eluded prior discovery not so much because it is rare or has an extremely restricted distribution, but because it simply lives somewhere that scientists have not yet been able to survey. Hence, there are usually few, if any, conservation implications in this context.
personal communication 2006
2.1. Criteria for determining sensitivity
The National Biodiversity Network (NBN) in the UK (Countryside Agencies OIN 2007), and the Department of Environment and Conservation in New South Wales, Australia (Department of Environment and Conservation 2007) developed detailed sensitivity criteria, and the previous version of this publication (Chapman and Oliver 2008) relied heavily on the work of those two agencies. Since the publication of the previous Guide, both these agencies (DECCW 2009, NBN 2019a, NBN 2019b, OEH 2019a) along with the South African National Biodiversity Institute (SANBI 2010, SANBI 2016), the Atlas of Living Australia (ALA 2018a) and others, have given a lot of thought to criteria for determining sensitivity within their jurisdictions. Documentation from all of them have contributed greatly to this document.
A series of criteria for determining the sensitivity of taxa and data along with recommended metadata statements for documenting the reasons for the determination are set out in Table 1. The first two are for use by biodiversity data holders and those creating trigger lists of potentially sensitive taxa and refer largely to the taxa themselves. The last two are for use by biodiversity data holders and deal with an assessment of the data they hold and are considering making available – they are not suitable for the creation of trigger lists.
The criteria are used to determine:
1. Risk of harm |
An assessment of whether the taxon is subject to harmful human activity. |
2. Impact of harm |
An assessment of the sensitivity of the taxon to the harmful human activity. |
3. Sensitivity of data |
An assessment on whether the release of data will increase harm. |
4. Decision on release and category of sensitivity |
A balanced decision regarding the release of the data and a determination of the category of sensitivity, and thus the level of generalization, of the data for release. |
A set of scenarios using Criteria 1 and 2 above to determine triggers for sensitivity of taxa is attached as an Annex to this document.
The first step in the process of determining sensitivity is to make an assessment on whether or not the taxon is subject to a harmful human activity or not and if the availability of related biodiversity data will increase the likelihood of the harmful activity occurring.
If it is not then there would appear no reason to list it as a potential environmentally sensitive taxon. It is recommended that you use the documented wording supplied but with additional supporting rational documenting the specifics of the threat, for example:
The taxon is at risk from harmful human activity – it is subject to attack by Phytophthora which is transported by human operated vehicles.
1.1. Is the taxon subject to a harmful human activity? |
|
YES |
NO |
1.2. Is there established evidence of current or recent occurrences of the harmful human activity? |
|
YES |
NO |
1.3. Will availability of related biodiversity data increase the likelihood of the harmful human activity taking place? |
|
YES |
NO |
The next step is to determine if the taxon is sensitive to that human harm or whether they are suitably robust not to be adversely affected.
2.1. Does the taxon have characteristics that make it significantly vulnerable to the harmful human activity? |
|
YES |
NO |
2.2. Is the taxon vulnerable to harmful human activity over its total range, or are there areas (such as in conservation zones, or other parts of the world) where the taxon is not at the same level of risk? |
|
YES |
NO |
Once it has been decided that the taxon is subject to a significant risk and impact from harm or not, then a decision needs to be taken on whether the release of specific data on that taxon – or other related data – will increase the risk and impact of harm.
3.1. Is the content and detail of the biodiversity data such that their release would enable someone to carry out a harmful activity upon the taxon or attribute? |
|
YES |
NO |
3.2. Is information already in the public domain, or already known to those individuals or groups likely to undertake the harmful activity? |
|
YES |
NO |
3.3. Would disclosure damage a partnership or relationship (especially where the maintenance of which is essential to helping achieve a specific conservation objective)? |
|
YES |
NO |
3.4. Would disclosure allow the locations of sensitive features to be derived through combination with other publicly available information sources? |
|
YES |
NO |
The final step is to make an overall assessment based on the three criteria above and to document the overall decision using the combined information documented in making each of the earlier decisions. Once it has been determined that the data should or should not be released, then it is important that a decision is made on the Category of sensitivity, and the level of generalization for the release of the data.
4.1. On balance, considering criteria 1 to 3 above and any important wider context, will withholding the information increase the risk of environmental harm or harm to a living person? |
|
YES |
NO |
4.2. Is the taxon distinctive and of high biological significance, under high threat from exploitation/disease or other identifiable threat where even general locality information may threaten the taxon? Or could the release of any part of the record cause irreparable harm to the environment or to an individual? |
|
YES |
NO |
4.3. Is the taxon such that the provision of precise locations at finer than 0.1 degrees (~10 km) would subject the taxon to threats such as disturbance and exploitation? Or does the record include highly sensitive information, the release of which could cause extreme harm to an individual or the environment? |
|
YES |
NO |
4.4. Is the taxon such that the provision of precise locations at finer than 0.01 degrees (~1 km) would subject the species to threats such as collection or deliberate damage? Or does the record include sensitive information, the release of which could cause harm to an individual or the environment? |
|
YES |
NO |
4.5. Is the taxon subject to low to medium threat if precise locations (i.e. locations with a precision greater than 0.001 degrees or 100m) become publicly available and where there is some risk of collection or deliberate damage? |
|
YES |
NO |
In the online survey (Chapman 2006), a number of respondents identified data awaiting publication, data subject to ongoing research, and incomplete or unchecked data as data that they would class as sensitive, and thus subject to restrictions on release. This is data whose sensitivity has a short time frame, and it is important that a time for release or review be clearly documented. They would most likely fall under criterion 3.3 above and would be documented accordingly with the supporting rationale being “awaiting publication”, etc.
| All data regarded as being sensitive should include a date for review of their sensitivity status, along with documented reasons for the sensitivity status. The date for review may be short or long depending on the nature of the sensitivity. |
The categories of sensitivity are largely based on those from the NSW Office of Environment and Heritage (DECCW 2009).
2.2. Categories of sensitivity
| Criterion | Reasoning |
|---|---|
|
Species or records for which no records will be provided at all, or which are only released as present within a large region such as a county, watershed, etc. |
The reason for non-disclosure is that:
Data may only be supplied under strict Licence conditions or as presence in a large region such as a watershed, county, or biogeographic region. |
|
Species or records for which coordinates will be publicly available ‘denatured’ (to 0.1 degrees) and/or other information in the record is generalized. Finer scale data (Category 3, Category 4 or detailed data) may be supplied to individuals under Licence. |
The reasons for restriction are that:
Data is supplied to the public
Data may be supplied at finer scales on request under the conditions of a written data agreement, usually a Data Licence Agreement. When data is provided to clients, they will be advised which species or fields are sensitive and may have their coordinates denatured to that available under Category 3 or Category 4. NB: In the case where the sensitivity is triggered by fields other than the georeference, it may be more appropriate to class the record as Category 3 or Category 4. |
|
Species or records for which coordinates will be publicly available ‘denatured’ (to 0.01 degrees) and/or other information in the record is generalized. Finer scale data (Category 4 or detailed data) may be supplied to individuals under Licence. |
The reasons for restriction are that:
Data is supplied to the public
Data may be supplied at finer scales on request under the conditions of a written data agreement, usually a Data Licence Agreement. When data is provided to clients, they will be advised which species or fields are sensitive and may have their coordinates denatured to that available under Category 4. NB: In the case where the sensitivity is triggered by fields other than the georeference, it may be more appropriate to class the record as Category 4. |
|
Species or records for which coordinates will be publicly available ‘denatured’ (to 0.001 degrees) and/or other information in the record is generalized. Detailed ‘as-held’ data may be supplied to individuals under Licence. |
The reasons for restriction are that:
Detailed data may be supplied under the conditions of a written data agreement, usually a Data Licence Agreement. When data is provided to clients, they will be advised which species or fields are sensitive. |
3. Generalizing textual information
In some cases, the information in text fields might be regarded as sensitive under certain circumstances. This may include such information as:
-
Names of living persons
-
Locality information
-
The date of collection
-
The collector’s number
-
Habitat
-
Landholder information
-
Taxonomic names
Some of these may need to be restricted to stop correlational analyses leading to deductions on the localities of records that are restricted or generalized – for example the collector’s name, date, and collector’s numbers in sequence. In other cases, it may be necessary to hide the name of a taxon in a list of collections in a biodiversity hotspot or sensitive locality.
Such restrictions should not restrict the provision of the record as a whole. The data that needs to be hidden may be removed and replaced with suitable wording (see below), or generalized–for example, just giving the name of a higher level taxonomic rank where the species is to be restricted.
| Whenever data in a textual field is restricted or generalized for distribution (such as the name of a collector, textual locality information, etc.), it should be documented by replacing it with appropriate wording-the field should not be left blank or null. |
Examples of replacement wording include:
name suppressed for reasons of privacy
This specimen represents an endangered or threatened species. The specific locality has been removed from the online record to protect this species from over-collection. These data may be supplied to researchers on request.
This specimen represents an endangered or threatened species. The specific locality has been generalized to presence within a grid of 0.1 degree resolution. Detailed data may be supplied to researchers on request.
| Where there is a need to restrict a taxonomic name (for example of sensitive taxa as part of a survey or checklist), it may be possible to replace it with a higher taxon name (genus/family, etc.) or to just report that there are ‘x’ sensitive taxa present without providing names. |
Occasionally, data providers may be tempted to restrict information in records related to a sensitive record (in addition to the sensitive record itself), such as the collector’s name and numbers in a sequence of records collected at the same location and time as a sensitive record in order to reduce the possibility of the sensitive record being found through correlational analysis. However, if the collector’s name and number is removed from just the sensitive record and not the others, it is unlikely that these could be deduced unless the seeker of the information already has inside knowledge. For this reason, and others, it is recommended that the data on related records not be restricted.
4. Generalizing spatial information
4.1. Generalization versus randomization
Few of the respondents to the online survey (Chapman 2006) recorded that they randomize data as opposed to generalizing it. Reasons for not randomizing included the extra work and computation involved, the increased chance of mistakes being made, and the less reliability that users may be able to place in the data.
Some respondents to the survey stated that they were comfortable with displaying presence/absence of sensitive data within large polygons or grids squares, because it still reflected the real data, but were aghast at the idea of deliberately ‘faking’ point coordinates such that locations appear as precise representations, but are randomly offset from the real data – i.e., they represent the deliberate introduction of error. Whereas generalization creates/retains ‘true’ data, randomization creates deliberately ‘false’ data.
In the fields of data mining and protection of privacy (including of individual privacy in census data), it is generally regarded that bottom-up generalization is far more practical and scientifically defensible than randomization (see for example, Wang et al. 2004, Dalvi and Keole 2015).
Another advantage of generalization is that it:
-
scales up, allowing the use of a consistent methodology at different scales
-
can be set to give different people different resolutions, depending on set roles, etc.
-
can simply provide for different scales of generalization for different categories of sensitivity
-
is easier to implement for those with low technological expertise
| Randomization is a methodology that I strongly recommend not be used. |
4.2. Generalization
One of the most common requirements for generalizing sensitive biodiversity information is to generalize the spatial locality or geographic coordinates (Chapman and Wieczorek 2020). Traditionally this has been done in many ways, and there has been little consistency in methodologies and very little documentation as to what has been done in each case. This has considerably reduced the value of the data for analysis, and often users are unaware that the data has even been modified.
Generalization (at least in a spatial sense) is usually of one of two types, namely:
-
Generalization to a grid (metric or geographic)
-
Generalization to a polygon (socio-political region, country, biogeographic region, watershed)
Many respondents to the survey (Chapman 2006) argued for the simplicity of generalization to a grid. The reasons given included:
-
the simplicity of being able to vary the scale for different categories of sensitivity,
-
the ease of maintenance and training, and
-
the simplicity of creating suitable documentation.
Generalizing to a grid, while protecting the exact locations of sensitive taxa, also provides data in a format that is still useable for a majority of users, especially where a standard grid is used.
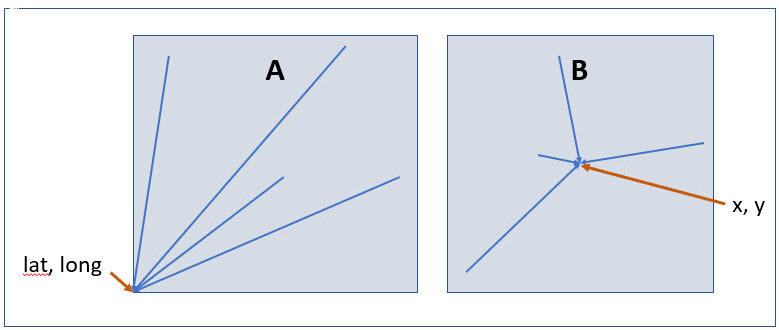
Figure 1 shows two methods that are commonly used to grid data. The first (recommended here) is a geographic grid (e.g. cartographic grids such as those based on geographic coordinates), the second-a metric grid (e.g. Europe Equal Area Grid 2001 – EPSG:19986). How each of these should be handled with respect to determining location and uncertainty, see Chapman and Wieczorek (2020). We recommend the use of a geographic grid, as discussed below, because of the ease of preparation and documentation, and because biological data being shared via the Darwin Core standard (Wieczorek et al. 2012) would need to be converted from a metric grid to a geographic grid before publication, with subsequent loss of precision. However, as noted in Chapman and Wieczorek (2020),
using the southwest corner as the coordinate for a point-radius georeference is wasteful, since the geographic radial would be from there to the farthest corner, which would be twice as far as it would be if the center of the grid cell was used instead. In any case, the characteristics of the grid should be recorded with the locality information.
Where data is generalized to a geographic or biogeographic region (a polygon), the data has less usability for many analyses, but was seen by many as a more secure way of ‘hiding’ sensitive data locations. Currently, GBIF.org has only limited ability to incorporate polygon data. There are some parallels with this method with the reporting of census results in many countries where summaries are reported using Statistical Local Areas or census tracts to restrict possible identification of individuals (Australian Bureau of Statistics 2006). A key difference is that with census data results are summarized over many individuals within a region, whereas with biological data we want to hide the location of a single entity within an area. It does de facto produce a summary, but this is not the primary intent. One problem with this method is that there is no guarantee that political (or even biogeographic) boundaries will remain constant over time and this further reduces the value of the data for many purposes. This has already been found to be a problem when comparing census data over time where census districts or tracts have altered. (Noble et al. 2011).
If georeferences are given for data that is generalized to a biogeographic or political region, the result can be quite misleading – a coastal species, for example, may end up with a georeference that is hundreds of kilometres inland, reducing usefulness for analysis or data cleaning. Making such data available without suitable documentation can lead to quite disastrous results for users. It is often better in some cases to not supply a georeference, but if one is supplied, then documentation should be clear with either a large Uncertainty Radius, or well documented Spatial Fit (see discussion under Spatial fit below).
Good practice dictates that whatever you do to generalize the data that you document it so that users of the data know what reliance they can place in it. Note, that when documenting what has been done, it is essential that both the coordinates and the coordinate uncertainty in meters be recorded. How this is best done can be seen in the Georeferencing Quick Reference Guide (Zermoglio et al. 2020).
I recommend that data providers who are generalizing their data do so using a standard methodology (see below), and to document this accordingly. As most biodiversity data is currently made available using decimal degrees (Chapman and Wieczorek 2020), the recommended method means that protocols such as Darwin Core (Wieczorek et al. 2012) do not need modification, other than to allow for suitable metadata documentation.
The method recommended below allows for several levels of generalization that conform to Categories 1–4 described in Table 1.
The recommended method for generalization is:
Category |
Sensitivity |
Georeference |
Category 1 |
Extreme |
Geographic coordinates not released or data may be released by watershed/bioregion/county, rounded to 1 degree, etc. |
Category 2 |
High |
Geographic coordinates rounded to 0.1 degree |
Category 3 |
Medium |
Geographic coordinates rounded to 0.01 degree |
Category 4 |
Low |
Geographic coordinates rounded to 0.001 degree |
Not sensitive |
Not sensitive |
Geographic coordinates unrestricted |
The South African National Biodiversity Institute, in largely implementing the criteria as laid out in the previous document (Chapman and Grafton 2008), have implemented only two categories (SANBI 2010, SANBI 2016): the original, non-generalized data and data generalized to one-quarter of a degree (0.25 degree (QDS)). We believe that this is a more difficult generalization to implement, and, unless fully and clearly documented, it could lead to a misleading level of precision. It also reduces the flexibility provided by the above four-categorization method.
4.3. Documentation
It is important to document the method and level of generalization so that users are aware of what has been done to the data, and what reliability they may be able to place in the data. Currently, neither Darwin Core nor the ABCD protocols provide fields for the recommended metadata. It has been proposed that these protocols be modified to accept such metadata (see Afterword), but in the meantime, it is recommended that the information be recorded using existing Darwin Core terms at the record-level (e.g., dwc:informationWithheld, dwc:dataGeneralizations or any of the 'Remarks' fields).
As far as the generalization of georeferencing data is concerned it is important to record that the data has been generalized using a ‘decimal geographic grid’ and record both:
-
Precision of the data provided (e.g. 0.1 degree; 0.001 degree, etc.)
-
Precision of the data stored or held (e.g. 0.0001 degree, 0.1 minute, 1 second, 100m square, etc.)
The recommendations for metadata for inclusion in the Darwin Core Location Class (TDWG 2018) are set out in the Afterword. Once they (or similar) have been adopted, then it is recommended that the appropriate fields be recorded and distributed with the data.
4.4. Duplicates and GUIDS
With plants, especially, and with other taxa (like insects), collectors often gather multiple specimens (duplicates or parts of sets)—usually on the order of four to six, though examples of more than 80 have been cited (Paul Morris 2007, personal communication, April)—with these duplicates or parts of sets often sent to many institutions around the world. One problem that arises is originating institutions may lose control of what happens to the information (including locality information) distributed to collections from those secondary institutions – remembering that the duplicates may have been distributed prior to the taxon being identified as sensitive.
In most cases this exchange of information is not a problem, but with sensitive taxa, it often is. The secondary institution may not know what are regarded as ‘sensitive taxa’ in the jurisdiction of the originating institution or may not have flagged that information. Sensitivity is not always information that can be distributed along with the collections, as it may not be known until much later that the species is endangered and/or sensitive. This issue is a difficult one, as simply labelling a taxon as sensitive may not be the answer: a taxon may be endangered in its native area (and thus sensitive) and may be a weed or pest in other areas, with locality information important for its control in both instances.
Identifying duplicates across institutions is not easy, as, especially for historic and legacy collections, it is often difficult to determine duplicate specimens. Some institutions, such as Centro de Referência em Informação Ambiental (CRIA) in Brazil in its speciesLink project and the Atlas of Living Australia, use matching across a number of fields such as collector number, date and locality, while GBIF is developing an algorithm for data-clustering. Currently, however, there is no universal global system available. The use of unique, persistent and resolvable Globally Unique Identifiers (GUIDs) (Page 2009, Richards 2010, Richards et al. 2011) will aid these processes in the longer-term, but the implementation of specimen-level GUIDs still seems some way off. A recent paper by Nelson et al. 2018 makes a number of recommendations on minting, managing and sharing GUIDs for herbarium specimens, but until such techniques are more widely adopted, identifying duplicates across institutions will remain an issue.
5. Documentation and metadata
It is important that data is accurately documented so that users and others know exactly what the data represents, and the reliance that can be placed in it. For example, a user needs the information to determine if the data is suitable for the analysis they are about to run. Many data providers reported in the survey (Chapman 2006) that one reason that they were reluctant to release some of their data was a fear that the data would be misused. If the data isn’t adequately documented, then the likelihood of inadvertent mis-use is greatly increased as the user may use the data in an analysis mistakenly thinking they are getting accurate point records, when in reality, the data had been generalized to a 10 km grid square, and could be anywhere in a 100 square kilometre area. If running a climate modelling algorithm, for example, then this sort of error could result in a quite misleading result. For this reason alone, it is important to data providers, data users, and end users (such as environmental managers, policy makers, etc.) that the data is accurately described.
In particular, there should be a clear documentation of the access constraints which could include, for example, an indication of which parts of the data is sensitive (if any), reasons for sensitivity and conditions under which release is possible.
5.1. Documenting sensitivity
Metadata fulfils an essential function regarding communication to third parties, of access constraints and use conditions that the data generators intend to give to their data. It can be considered as an ‘aid’ in protecting data and information, since it will allow system users to visualize the conditions established by the data generator for access and use of the information. Additionally, in case the data are not accessible, the metadata allows knowledge of the conditions of access through other media (digital or not) as well as a summary of the content. (Llinás 2005)
Metadata has generally been used to refer to documentation of a whole dataset. Documentation at the record level has usually been referred to just in comments. I prefer, however, to term this ‘record-level metadata’, and to formalize the process. In the previous chapter a recommendation was made that where data is generalized for distribution, to document the level of generalization – for example, that the data had been generalized using a decimal geographic grid, and to record both the precision of the data provided and the precision of the data ‘as-held’ or stored. Also, in the chapter on Determining sensitivity, a series of documentation processes were recommended (Table 1). Some of these may be more appropriate for documenting the reasons for regarding a taxon as a potential environmentally sensitivity taxon (Criteria 1 and 2), while the others (Criteria 3 and 4) are appropriate to the data themselves and belong as part of the broader record-level metadata. To fully document the reasons for restricting data, however, it may be necessary to inherit the documentation from Criteria 1 and 2 to the record level – for example, the reason that data is restricted may include that the taxon is subject to harmful human activity.
At the moment, neither the Darwin Core nor the ABCD standards have fields for recording the type of record-level metadata that is recommended here. A number of recommendations have been made to Biodiversity Information Standards (TDWG) for the inclusion of extra fields to the Darwin Core Location Class (TDWG 2018) and are listed in the Afterword.
Until such time as these standards and protocols are modified, it is recommended that the data be documented using existing Darwin Core terms at the record-level (e.g., dwc:informationWithheld, dwc:dataGeneralizations or any of the 'Remarks' fields), and, as far as possible, to record the same type of information that would be included in the recommended fields, for example, that:
-
The data is sensitive
-
The primary reasons the data is regarded as sensitive (see Criteria 1–4, Table 1) along with supporting rationale
-
The date that the sensitivity of the data should be reviewed
-
Precision of the data made available
-
Precision of the original data stored or retained
When noting access constraints in the metadata at the dataset level, you may include something like:
This dataset is only available to the public at a summary resolution for the following reason. Some of the information held within this dataset relates to species that are vulnerable to human disturbance or prejudice. Two species (Adelanthus lindenbergianus, Athalamia hyaline) are significantly vulnerable to collecting. The full detail of this sensitive information may be made available under licence to specific organizations and individuals that need to know to avoid harm to the environment. Please contact the provider for more information.
5.2. Spatial fit
Spatial Fit (Chapman and Wieczorek 2006, 2020) is a concept that has arisen out of the BioGeomancer project (Guralnick et al. 2006) and provides a measure of how well a geometric representation matches the original spatial representation. Spatial fit is a value of either zero, one or greater than 1, where 1 represents an exact match (i.e. the data has not been generalized). Details on how Spatial Fit may be calculated can be found in Chapman and Wieczorek (2006, 2020), where the summary below can also be found:
A spatial fit with a value of 1 is an exact match or 100% overlap. If the geometry given does not completely encompass the original spatial representation, then the spatial fit is zero (i.e., some of the original is outside the transformed version, which we interpret as not being a fit). If the transformed shape does completely encompass the original spatial representation, then the value of the spatial fit is the ratio of the area of the transformed geometry to the area of the original spatial representation. Special case: If the original spatial representation is a point and the geometry presented in not a point, then the spatial fit is undefined.
With respect to generalization of data, the Spatial Fit can be seen as a Generalization Coefficient – i.e. to what extent a record has been generalized – a ratio of the generalized area to the true area.
Spatial Fit = Generalized area/True area
An example of its applicability is where a georeference with an uncertainty radius of 1 km (using a point radius method (Wieczorek et al. 2004)) is made available using a 10 km² grid (which completely covers the uncertainty). In this case the Spatial Fit would be greater than 1 as it represents an area greater than the real uncertainty (as shown in Figure 2).
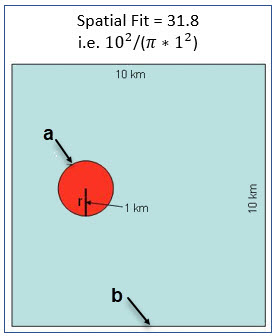
The smaller the grid size, the closer the Spatial Fit will be to ‘1’. Note that a record that has its georeference randomized or generalized such that a portion of the uncertainty radius falls outside the grid square would have a Spatial Fit equal to zero.
6. Authentication and Authorization
As recommended by the experts’ workshop, and identified by many in the online survey (Chapman 2006), responsibility for determining who may or may not have access to detailed data on sensitive data, possibly through the use of secure log-on, or one-off data licence agreements, must be with the data providers.
It was also agreed at the workshop that it is not the role of GBIF to manage the identification, verification or authorization of users, nor to control authentication or log-on at the Data Portal, but it may have a role in providing guidance and a suitable authentication method to the Nodes.
It was reported at the experts’ workshop that the technical issues relating to the authentication of a group or individual, and the use of roles, etc. is not a difficult task. There are several well established protocols and working systems for authentication in use and these could easily be adapted for use by data providers.
The main issue is in determining who the authorized users should be and how to determine who are bona fide users and who are not. This is a difficult issue and one that will need to be explored over time. It is not something that can be recommended in this best practices document; however the earlier report (Chapman 2007) did make a number of recommendations on how this issue may be further explored.
7. Implementations
Since the publication of the Guide (Chapman and Grafton 2008), several countries, state and local jurisdictions have developed their own sensitive taxa data policies. Jurisdictions often have individual circumstances (legal, ethical, practical, etc.) that may dictate the way their own policies need to be implemented. In developing your own policies, in addition to relying on this document, you may wish to look at those jurisdictional policies. I would caution, however, in diverting too far away from the criteria and principles laid out in this document, due to the complications that may arise in sharing data, especially through aggregating and publishing initiatives such as GBIF. The main purpose of this document is to encourage the adoption and use of standardized methodologies. Most of the jurisdictional documents will, of necessity, go further than this document which is only based around generalization. This document does not go into other aspects such as the privacy of individuals, licensing, etc. Any derived works should be made publicly accessible and clearly lay out the policies with respect to the publication and sharing of data. Where policies differ to those documented here, those differences should be noted along with explanations as to the reasons for differing.
The South African National Biodiversity Institute (SANBI) has listed four types of data on sensitive taxa that may need to be restricted (SANBI 2010):
-
Data on population sizes or numbers of a sensitive taxon or its populations that might influence the rarity value or commercial value of the taxon;
-
Data about the habitat and ecosystem of a sensitive taxon that may allow the locality of the taxon to be inferred;
-
Geo-referenced data about a sensitive taxon (including precise locational data, descriptions of locations and/or localities and point locality coordinates) that may allow populations to be located;
-
Records of specimens in collections or observation records (including the name of the collector, the collector specimen number, taxon identity, the locality description, coordinates more precise than a quarter-degree-square, population size, date of collection, collector of the specimen, and any habitat information associated with the specimen), that with analysis may allow a population to be located. The record may refer to a single specimen, or a sample, which includes several or many specimens with identical collecting details. All specimen/observation records for a sensitive taxon, no matter when collected, would be equally restricted.
SANBI have not adopted the categorizations recommended here, adopting just the one level of generalization at what they call QDS (Quarter Degree Square) or about 25 km by 25 km (see discussion in SANBI 2016).
The Atlas of Living Australia (ALA) has developed a policy that places sensitive taxa into three categories (ALA 2018a):
-
conservation, e.g. the locations of threatened species
-
biosecurity, e.g. unverified sightings of pests not previously recorded in Australia
-
privacy, e.g. specimens collected on private property, collectors names, etc.
This policy is based on a set of guiding principles (Tann and Flemons 2009):
-
making scientific data readily accessible
-
minimizing harm by explicitly restricting access to selected sensitive information
-
assisting State and federal authorities with their obligations under Freedom of Information
-
assisting State and federal authorities with Australia’s international trade obligations
-
assisting with the sharing of data held in trust by a custodian, where there is an agreement or expectation that this data will not be misused
-
respect for the differences of approach to sensitive data in jurisdictions across Australia
-
respect for privacy and restrictions to personal information
Unfortunately, to date, each Australian State has different individual policies on what taxa are regarded as sensitive, and on how data on sensitive taxa are shared. Discussion on the development of a national list of sensitive taxa is ongoing, but agreement on a consistent policy for sharing that data still seems to be some way off.
The Australian Department of the Environment (Australian Government 2016) have adopted the levels of generalization as set out in this document and the policy document provides a number of well worked examples. The policy includes some additional guiding principles including:
-
Be accountable: The decision to restrict access to data needs to be justifiable, consistent and repeatable and abide by relevant legislation, regulations or policy.
-
Decisions made closest to source: The data custodian should have responsibility for determining whether ecological data should be classified as sensitive.
-
Retain the original data: Data custodians must retain an unaltered original version of the ecological data and safeguard this original version.
-
Transparency: Documentation should be linked to the data and must be available to all users of the data. Documentation ensures potential data users understand what data exists, why it was classed as sensitive and how it has been altered or protected.
-
Respect dataset restrictions: Data custodians should not release data that has not been processed in accordance with this policy
-
Review over time: Data custodians should do regular reviews (every 2–5 years) on datasets to determine if their context has changed. What is currently considered sensitive data may not be sensitive in the future.
One principle that is emphasized by the policy is that if data on the identified sensitivities (e.g. location) is already publicly available, then it is unlikely that data can be considered sensitive.
The New South Wales Office of Environment and Heritage (DECCW 2007, OEH 2019a) notes that “some threatened species are very sensitive to disturbance and exploitation. Information about the location of these species is considered ‘sensitive’ and OEH will not provide it to third parties, with some limited exceptions. Precise locational information about sensitive species is exempt from freedom of information requests.” Their sensitive data policy has three categorized levels:
Category 1: Species of high biological significance, for which no records will be provided at all. The reason for non-disclosure is that the species is highly threatened by exploitation/disease or other identifiable threats, and even general locality information may threaten the taxon. The famed Wollemi pine falls into this category.
Category 2: Species considered to be at serious risk from threats such as disturbance or exploitation. For species in this category, geographic coordinates of sightings will be supplied ‘denatured’, in order to generalize the locality. Exceptions to this rule may be granted to some government agencies, or for certain research purposes.
Category 3: Species considered to be at medium to high risk of threats such as disturbance or exploitation. For species in this category, coordinates will be supplied at ’as held’ accuracy to licensed clients, but will otherwise be supplied ‘denatured’.
They include 3 basic categories of ‘denaturing’ (OEH 2011) that are largely consistent with what is recommended here:
-
Category 1: no records provided
-
Category 2:
-
records denatured to 0.1° (~10 km) for public web applications
-
records denatured to 0.01° (~1 km) for provision to licensed clients
-
-
Category 3:
-
records denatured to 0.01° (~10 km) for public web applications
-
records provided ‘as held’ to licensed clients
-
A list of sensitive species, the category of sensitivity, and the reasons why they are regarded as sensitive is maintained and published on the agency’s website (OEH 2019a).
The UK National Biodiversity Network (NBN 2019b) uses criteria that allows for different categories of generalization in the different member countries of the UK (e.g. a species can be listed as sensitive in Wales, but not in Scotland). Records are submitted to the NBN Atlas at the best capture resolution. The location of sensitive species are generalized and the generalized data made available to the public under a Creative Commons licence as determined by the data provider. Lists of sensitive taxa, along with the reasons for sensitivity, and the generalization level for each of England, Scotland and Wales are maintained and published.
Birdlife Australia (Birdata) have developed a Sensitive Species policy based on the principles and generalization categories as set out in this and the previous Guide (Chapman and Grafton 2008). They have identified six categories of data where the localities may need to be generalized:
-
Wildlife poaching and international trade
-
Legal and illegal hunters including trophy, recreational, commercial and sport hunters
-
Egg collectors
-
Illegal capture of wild birds for the cage trade and falconry
-
Wildlife enthusiasts exhibiting intrusive behaviour, particularly to territorial species
-
Trespassing/accessing private property or indigenous protected areas without a permit.
The US Forest Service has a policy for sensitive species to ensure viable populations throughout their geographic ranges. Once the objectives are accomplished and viability is no longer a concern, species shall not have “sensitive” status (US Forest Service 2005). Sensitive species are those plant and animal species identified by the Regional Forester for which population viability is a concern on National Forest Service (NFS) lands within the region. The goal of the Forest Service Sensitive Species Program is to ensure that species numbers and population distribution are adequate so that no federal listing will be required and no extirpation will occur on NFS lands (US Forest Service 2016).
No specific mention is made of different categories, or of generalizing location information for the public. However, according to Hartter et al. (2013), the US Forest Service seeks to protect research sites by not disclosing geospatial references along with its data.
Natural Resources Canada and GeoConnections Canada commissioned a study to develop Best Practices for Sharing Sensitive Environmental Geospatial Data (AMEC Earth and Environmental 2010). The Guidelines consider environmental geospatial data to be “thematic geospatial data that could be used for analysis in areas such as environmental impact assessments, land use planning, land management, sustainable development, resource management, airshed management, etc.” The document lists five criteria for determining sensitivity. The third criterion includes the data considered in this document:
Natural Resource Protection: the use of the information can result in the degradation of an environmentally significant site or resource
The document recommends that as Canada is a member of GBIF, Canadian organizations, should incorporate the Guide to Best Practices for Generalising Sensitive Species Occurrence Data (Chapman and Grafton 2008) when assessing their environmental datasets. Without mentioning specific generalization levels, the document does site the categories of generalization in this and the previous Guide.
Other aggregation agencies, such as iDigBio, have left it to those supplying the data to deal with sensitivity, and have not developed a policy per se.
“iDigBio accepts all Data it receives via the Services as-is. It makes no effort to mask Sensitive Data. The Data Publisher is solely responsible to mask or withhold information, including Sensitive Data, from the public.”
In many cases, decisions on whether to release data to the public is done on a project to project basis. For example, Fong and Qiao (2010) describe a project to map locations of an endangered species of turtle in China and argue that while this location data is valuable to researchers, it should not be made publicly available due to concerns about the safety of the animals.
Afterword
Listing sensitive taxa
Data is already distributed around the globe through duplicate specimens, etc., and although data may be restricted from some institutions, others holding duplicates may be releasing the same information. This may be through ignorance of what may be regarded as sensitive in the home ranges of the taxon concerned as no universal list of what is regarded as ‘sensitive’ is currently available. Difficulties are compounded by the fact that a taxon may be sensitive in one area, but not in another (and indeed may even be an invasive weed or pest species in the second location). If identical data is publicly available through other sources, it cannot be considered sensitive (Australian Government 2016).
For these reasons, it has been recommended that a trigger list of potential environmentally sensitive taxa should be created and linked through GBIF’s Backbone Taxonomy. This would have the advantages of alerting data providers in other jurisdictions that a species is potentially sensitive, and via the Backbone Taxonomy would provide links to synonyms. It is important to note that the list should be regarded as a trigger to flag the need for a decision on the actual sensitivity of sharing information using the criteria in the previous chapter, and not for generating blanket restrictions. Not all endangered species are threatened through knowledge of their locations, or across the totality of their range, and so should not be regarded as sensitive per se and thus the list of potential environmentally sensitive taxa should be much smaller than any existing list of rare and threatened species.
The list should be created using Criteria 1 and 2 and scenarios in Annex 1 and include additional information, such as:
-
Name of taxon
-
Criteria and supporting rationale for inclusion
-
Name of person or organization responsible for the taxon being included
-
Geographic coverage of sensitivity (especially if only sensitive over part of its range or within one jurisdiction)
-
Recommended sensitivity category
-
Date for review
Jurisdictions may also wish to maintain a similar list for their own purposes, and it is recommended that if they do so, they include the above information in all cases. The advantages of making the information more broadly available is that it will alert other data custodians that your jurisdiction regards the taxon as potentially sensitive, and alert users that they should take the sensitivity into account when publishing the results of their analyses, etc.
| Any list of potential environmentally sensitive taxa should be regarded as a trigger only and any restrictions on the availability of actual data should be made on a case by case basis taking into account the listed criteria. |
Metadata recommendations
A number of recommendations have been made to Biodiversity Information Standards (TDWG) for the inclusion of extra fields to the Darwin Core Location Class (TDWG 2018). The recommendations included:
Field |
Comments |
dataSensitiveIndicator |
Y/N flag that the observation is sensitive. |
dataSensitiveReason |
The primary reason why the data is sensitive. Suggested format is either a picklist with values derived from Criteria 1–4 above (or a text field that combines the statements 1a–4g attached to those criteria). |
dataSensitiveComments |
Further free-text information on the reason(s) or supporting rationale for determining relevance of the Criteria for this record as recommended above. |
sensitiveDateForReview |
A date field documenting when the sensitive nature of the date should be reviewed. Especially important where the sensitivity is just awaiting publication of results, etc. |
precisionDataProvided |
The scale or the precision of the data made available via the Darwin Core record – may be done as coordinate precision, e.g.
|
precisionDataStored |
The scale or the precision of the data stored or retained by the data custodian – may be done as coordinate precision, e.g.
or maybe more free text – such as ‘1 minute’, ‘0.1 minute’, ‘1 second’ – depending on how data is stored. |
Glossary
- authentication
-
refers to the determination of a user’s identity, as well as determining what a user is authorized to access. The most common form of authentication is user-name and password, although this also provides the lowest level of security.
- authorization
-
refers to the process of determining which individuals can be afforded different access rights for authentication and data access.
- decimal degrees
-
degrees expressed as a single real number (e.g. −22.343456). Note that latitudes south of the equator are negative, as are longitudes west of the prime meridian to −180 degrees.
- generalization
-
refers here to any modifications carried out to source data to conceal sensitive content, typically by reducing the precision of the data (such as reporting at the level of a watershed, grid or county, citing just the nearest named place, or by deleting some parts of the data). In geographic terms it refers to the conversion of a geographic representation to one with less resolution and less information content; traditionally associated with a change in scale. Also referred elsewhere to as: fuzzying, dummying-up, etc.
- geographic coordinates
-
a measurement of a location on the earth’s surface expressed in degrees of latitude and longitude.
- harmful human activity
-
human activities or processes that have had, are having or may have an adverse impact on the status of the taxon under assessed. Examples include unsustainable fishing or logging, hunting, harvesting, agriculture, housing developments, among others (see IUCN 2020).
- precision
-
describes the finest unit of measurement used to express a value (e.g. if a record is reported to the nearest second, the precision is 1/3600th of a degree; if a decimal degree is reported to two decimal places, the precision is 0.01 of a degree).
- randomization
-
refers to a deliberate haphazard arrangement of observations so as to obscure their true location. Randomization leads to a falsification of the data. Also referred to as falsifying.
- record-level metadata
-
refers to documentation at the level of a record rather than for a complete dataset. In this document it largely refers to documentation of the sensitivity status of the record (or the species of which it is a part) along with access constraints pertaining to the record and details of any generalization of the data.
- sensitive data
-
any data, that because of their nature, a data provider does not want to make available in their raw state, e.g. precise localities of endangered taxa.
- spatial fit
-
a measure of how well one geometric representation matches another geometric representation as a ratio of the area of the larger of the two to the area of the smaller one (see Figure 2) (Chapman and Wieczorek 2006 & Chapman and Wieczorek 2020).
Acknowledgements
The earlier document (Chapman and Grafton 2008) acknowledged the many people who responded to the initial survey and who attended the workshops and who were valuable in the preparation of that document.
In addition, I would now like to thank John Wieczorek and Paula Zermoglio, for the on-going support and advice, as well as staff at the GBIF Secretariat, especially Laura Russell and Kyle Copas who oversaw the project.
References
-
ALA (2018a) What is Sensitive Data? Atlas of Living Australia. https://support.ala.org.au/support/solutions/articles/6000195500-what-is-sensitive-data-
-
ALA (2018b) What is data licensing? Atlas of Living Australia. https://support.ala.org.au/support/solutions/articles/6000195495-what-is-data-licensing-
-
AMEC Earth & Environmental (2010) Best practices for sharing sensitive environmental geospatial data. Version 1.0 http://publications.gc.ca/collections/collection_2011/rncan-nrcan/M104-4-2010-eng.pdf
-
Australian Bureau of Statistics (2006) Statistical Local Area (SLA) in 2901.0 - Census Dictionary, 2006 (Reissue). https://www.abs.gov.au/AUSSTATS/abs@.nsf/bb8db737e2af84b8ca2571780015701e/23d04985e1786824ca25720b0002bb18!OpenDocument
-
Australian Government. Department of the Environment (2016) Sensitive Ecological Data - Access and Management Policy V1.0. https://www.environment.gov.au/system/files/resources/246e674a-feb1-4399-a678-be9f4b6a6800/files/sensitive-ecological-data-access-mgt-policy.pdf
-
Birdata (n.d.) Sensitive Species. Birdlife Australia. https://birdata.birdlife.org.au/sensitive-species
-
Chapman AD (2006) Questionnaire on Dealing with Sensitive Primary Species Occurrence Data: Summary of responses. Copenhagen: GBIF Secretariat. https://doi.org/10.35035/vs84-0p13
-
Chapman AD (2007) Dealing with Sensitive Primary Species Occurrence Data. Report. Report to the Global Biodiversity Information Facility. Copenhagen: GBIF Secretariat. https://doi.org/10.35035/rajc-t668
-
Chapman AD & Grafton O (2008) Guide to Best Practices for Generalising Sensitive Species Occurrence Data, version 1.0. Copenhagen: GBIF Secretariat. https://doi.org/10.15468/doc-b02j-gt10
-
Chapman AD & Wieczorek J, eds. (2006) Guide to Best Practices for Georeferencing. Copenhagen: GBIF Secretariat. https://doi.org/10.15468/doc-2zpf-zf42
-
Chapman AD & Wieczorek J (2020) Georeferencing Best Practices. Copenhagen: GBIF Secretariat. https://doi.org/10.15468/doc-gg7h-s853
-
Corti L, Day A, & Backhouse G (2000). Confidentiality and informed consent: Issues for consideration in the preservation of and provision of access to qualitative data archives. In Forum Qualitative Sozialforschung/Forum: Qualitative Social Research (Vol. 1). http://www.qualitative-research.net/index.php/fqs/article/viewArticle/1024 [Accessed 15 May 2019].
-
Countryside Agencies’ OIN (2007) The ‘Environmental Exception’ and access to information on sensitive features. Version 1.3.3, Countryside Agencies’ Open Information Network Environmental Information Regulations Guidance Note No. 1.
-
Dalvi VB & Keole RR (2015) Bottom-Up Generalization: A Data Mining Solution to Privacy Protection. International Journal of Science and Research 4(1): 2068–2071. https://www.ijsr.net/archive/v4i1/SUB15769.pdf
-
Department of Environment and Conservation − NSW (2007) Threatened Species Information Disclosure Policy (Version 3 Amended March 2007). No longer available: replaced by OEH 2019.
-
DECCW (2009) Sensitive Species Data Policy. New South Wales Department of Environment, Climate Change and Water. https://www.environment.nsw.gov.au/resources/nature/SensitiveSpeciesPolicyDEC09.pdf
-
Fong JJ & Qiao G (2010) New localities of endangered Chinese turtles from museum specimens and the practical and ethical challenges using and reporting natural history collection data. Zootaxa 2393: 59–68. https://doi.org/10.11646/zootaxa.2393.1.5
-
Frank D, Kriesberg A, Yakel E & Faniel IM (2015) Looting Hoards of Gold and Poaching Spotted Owls: Data Confidentiality Among Archaeologists and Zoologists. https://deepblue.lib.umich.edu/handle/2027.42/115883
-
GBIF Secretariat (2017) Data publisher agreement. https://www.gbif.org/en/terms/data-publisher
-
GBIF Secretariat (2019). GBIF Backbone Taxonomy. Checklist dataset https://doi.org/10.15468/39omei
-
Guralnick R, Wieczorek J, Beaman R, Hijmans RJ & the BioGeomancer Working Group (2006) BioGeomancer: Automated Georeferencing to Map the World’s Biodiversity Data. PLoS Biol 4(11): e381. https://doi.org/10.1371/journal.pbio.0040381
-
Guterman L (2006) Endangered by Research: Poachers mine the scientific literature for the locations of newly discovered animals. The Chronicle of Higher Education 52(46): A12. https://www.chronicle.com/article/Endangered-by-Research/26117
-
Hartter J, Ryan SJ, MacKenzie CA, Parker JN & Strasser CA (2013) Spatially Explicit Data: Stewardship and Ethical Challenges in Science. PLoS Biol. 11(9), e1001634. http://doi.org/10.1371/journal.pbio.1001634
-
IUCN (2020) Threats Classification Scheme, version 3.2. https://www.iucnredlist.org/resources/threat-classification-scheme
-
Llinás JV (2005) Data and Information on Biodiversity and its Protection in the Digital Realm Ver. 1. Bogotá, Colombia: Biological Resources Research Institute Alexandre von Humboldt.
-
NBN (2019a) The NBN Data Exchange Principles and their rational. National Biodiversity Network. https://nbn.org.uk/the-national-biodiversity-network/archive-information/data-exchange-principles/
-
NBN (2019b) Sensitive Data. National Biodiversity Network. https://nbn.org.uk/sensitive-data/
-
Nelson G, Sweeney P & Gilbert S (2018) Use of globally unique identifiers (GUIDs) to link herbarium specimen records to physical specimens. Applications in Plant Sciences 6(2): e1027. https://doi.org/10.1002/aps3.1027
-
Noble P, van Riper D, Ruggles S, Schroeder J & Hindman M (2011) Harmonizing Disparate Data across Time and Place: The Integrated Spatio-Temporal Aggregate Data Series. Historical Methods 44(2): 79-85. https://doi.org/10.1080/01615440.2011.563228
-
OEH (2011) Appendix 2. Denaturing Specifications for Sensitive Species records. Sydney, Australia: NSW Government. Office of Environment and Heritage. https://www.environment.nsw.gov.au/-/media/OEH/Corporate-Site/Documents/Animals-and-plants/Wildlife-management/appendix-2-denaturing-specifications-sensitive-species-records.pdf
-
OEH (2019a) Sensitive species data policy. Sydney, Australia: NSW Government. Office of Environment and Heritage. https://www.environment.nsw.gov.au/topics/animals-and-plants/wildlife-management/wildlife-policies-and-guidelines/sensitive-species-data [Accessed 26 May 2019].
-
OEH (2019b). Scientific Licenses. Sydney, Australia: NSW Government. Office of Environment and Heritage. https://www.environment.nsw.gov.au/licences-and-permits/scientific-licences
-
Page RDM (2009) bioGUID: Resolving, discovering, and minting identifiers for biodiversity informatics. BMC Bioinformatics 10(Suppl 14): S5. https://doi.org/10.1186/1471-2105-10-S14-S5
-
Parry O & Mauthner NS (2004) Whose Data are They Anyway?: Practical, Legal and Ethical Issues in Archiving Qualitative Research Data. Sociology 38(1): 139–152. https://doi.org/10.1177/0038038504039366
-
Richards K (2010) TDWG GUID applicability statement, version 2010‐09. Biodiversity Information Standards (TDWG). http://www.tdwg.org/standards/150
-
Richards K, White R, Nicolson N & Pyle R (2011) Beginners’ guide to persistent identifiers, version 1.0. Copenhagen: Global Biodiversity Information Facility. https://www.gbif.org/document/80575
-
SANBI (2010) Biodiversity Information Policy Framework. Policy Series. Digital Access to Sensitive Taxa Data. Silverton, South Africa: South African National Biodiversity Institute. http://biodiversityadvisor.sanbi.org/wp-content/uploads/2012/09/SANBI-Biodiversity-Information-Policy-Series-Digital-Access-to-Sensitive-Taxon.pdf
-
SANBI (2016) Report of the National Sensitive Species List Workshop 18 and 19 August 2016. Silverton, South Africa: South African National Biodiversity Institute. http://biodiversityadvisor.sanbi.org/wp-content/uploads/2017/06/20160819-NSSL-Workshop-Report.pdf
-
Stuart BL, Rhodin GJ, Grismer LL & Hansel T (2006) Scientific Description Can Imperil Species. Science 312(5777): 1137. https://doi.org/10.1126/science.312.5777.1137b
-
Tann J & Flemons P (2009) Our secrets are not our secrets. Atlas of Living Australia Sensitive Data Report. Version 1.1. https://www.ala.org.au/wp-content/uploads/2010/07/ALA-sensitive-data-report-and-proposed-policy-v1.1.pdf [Accessed 25 May 2019]
-
TDWG (2018) Darwin Core quick reference guide. Biodiversity Information Standards (TDWG). https://dwc.tdwg.org/terms/
-
US Forest Service (2005) Forest Service Sensitive Species Summary, Designated Sensitive Species (that are not listed or proposed species under the ESA), as of 31 October, 2005. https://www.fs.fed.us/biology/resources/pubs/tes/ss_sum_by_region_31Oct2005_fs.pdf
-
US Forest Service (2016) Forest Service Sensitive Species – Wildlife. https://www.fs.usda.gov/Internet/FSE_DOCUMENTS/fseprd530660.pdf
-
Wang K, Yu PS & Chakraborty S (2004) Bottom-Up Generalization: A Data Mining Solution to Privacy Protection, in Proceedings of Fourth International IEEE Conference on Data Mining (ICDM’04): 249-256.
-
Wang Z, Dong H, Kelly M, Macklin JA, Morris PJ, Morris R 2009. Filtered-Push: A Map-Reduce Platform for Collaborative Taxonomic Data Management. World Congress on Computer Science and Information Engineering, March 31 – April 2, 2009, Los Angeles, California, USA. https://doi.org/10.1109/CSIE.2009.948
-
Wieczorek J, Guo Q & Hijmans R (2004) The point-radius method for georeferencing locality descriptions and calculating associated uncertainty. International Journal of Geographical Information Science 18: 745-767.
-
Wieczorek J, Bloom D, Guralnick R, Blum S, Döring M, Giovanni R, Robertson T & Vieglais D (2012) Darwin Core: An Evolving Community-Developed Biodiversity Data Standard. PLoS ONE 7(1): e29715. https://doi.org/10.1371/journal.pone.0029715
-
Wylie A (1996) Ethical dilemmas in archaeological practice: Looting, repatriation, stewardship, and the (trans) formation of disciplinary identity. Perspectives on Science 4(2): 154–194.
-
Zermoglio PFF, Chapman AD, Wieczorek JR, Luna MC, Bloom DA (2020) Georeferencing Quick References Guide Copenhagen: Copenhagen: GBIF Secretariat. https://doi.org/10.35035/e09p-h128.
Annex 1: Scenarios using Criteria 1 and 2 as Triggers
The following sets of scenarios show how the criteria statements given in the Chapter on Determining sensitivity may be used to develop summary statements for documenting the reasons why a taxon may be regarded as sensitive. Summary statements should also include supporting rationale, such as specific types of harm, etc. For example, in scenario B, the full statement may read something like:
Taxa could be at risk from harm from disease carried on the wheels of forestry machinery but occurrence is not affected by data availability.
This may apply to a species of plant in a forestry area susceptible to Phytophthora attack, the fungi being transferred on the wheels of forestry vehicles.
Criterion 1
| Scenario A |
|---|
1a: There is no significant risk of a harmful human activity. |
The taxon is not sensitive. |
| Scenario B |
|---|
1a: The taxon is at risk from harmful human activity. |
1d: There is currently no established evidence of actual harm to the taxon. |
1f: Availability of biodiversity data will not increase the likelihood of the harmful human activity taking place. |
The taxon could be at risk from harm but likelihood of harm is not affected by data availability. |
| Scenario C |
|---|
1a: The taxon is at risk from harmful human activity. |
1d: There is currently no established evidence of actual harm to the taxon. |
1e: Availability of biodiversity data will increase the likelihood of the harmful human activity taking place. |
The taxon could be at risk from harm and the likelihood of harm is affected by data availability. |
| Scenario D |
|---|
1a: The taxon is at risk from harmful human activity. |
1c: There is established evidence of actual or recent harm to the taxon. |
1f: Availability of biodiversity data will not increase the likelihood of the harmful human activity taking place. |
The taxon is at risk from harm and there is evidence to support this, but occurrence is not affected by data availability. |
| Scenario E |
|---|
1a: The taxon is at risk from harmful human activity. |
1c: There is established evidence of actual harm to the taxon. |
1e: Availability of biodiversity data will increase the likelihood of the harmful human activity taking place. |
The taxon is at risk from harm, there is evidence to support this, and occurrence is affected by data availability. |
Criterion 2
| Scenario F |
|---|
2b: The taxon is not significantly vulnerable to the harmful human activity. |
2d: The taxon is not vulnerable to harmful human activity over its total range and/or there are areas where the taxon is not at significant risk. |
The taxon is not significantly vulnerable to the harmful activity, and is not vulnerable to that activity over its total range and there are areas where the taxon is not at significant risk from that activity. |
| Scenario G |
|---|
2a: The taxon has characteristics that make it significantly vulnerable to the harmful human activity. |
2d: The taxon is not vulnerable to harmful human activity over its total range and/or there are areas where the taxon is not at significant risk. |
The taxon is significantly vulnerable to the harmful activity, but is not vulnerable to that activity over its total range and there are areas where the taxon is not at significant risk from that activity. |
| Scenario H |
|---|
2a: The taxon has characteristics that make it significantly vulnerable to the harmful human activity. |
2c: The taxon is vulnerable to harmful human activity over its total range. |
The taxon is significantly vulnerable to the harmful activity, and is vulnerable to that activity over its total range. |